ParaView is deployed on ASC Purple "out-of-the-box" despite the complexity of hardware, and ParaView's quality parallel rendering and image delivery mechanism make remotely interacting with the data simple and effective. ParaView provides the remote analysis capability our scientists need. Any communication between these two sites required the use of SecureNET, an encrypted communication channel with moderate communication speeds (45-600 Mbps). of parallel visualization relied on parallel I/O and parallel rendering. HEDP simulations ran on the ASC Purple supercomputer located in Lawrence Livermore National Laboratory, over 850 miles (1300 km) from the analysts' office. ParaView was chosen to carry out the project and conduct the render performance. ParaView itself can’t render on multiple cores. So here’s a quick’n’dirty tutorial for dummies. Rendering results in ParaView on multiple cores on a desktop computer is a little less explained on the light side of the internets. Such was the case for a set of analysts and Sandia National Laboratory studying high energy density physics (HEDP). Using OpenFOAM in parallel with MPI is well explained in the OpenFOAM manual. Furthermore, the hardware resources may be located far away and without the high speed connections expected in a local network. The simulation hardware is often unique, complex, and built without visualization or post processing in mind. Moving such large quantities of data is infeasible. 
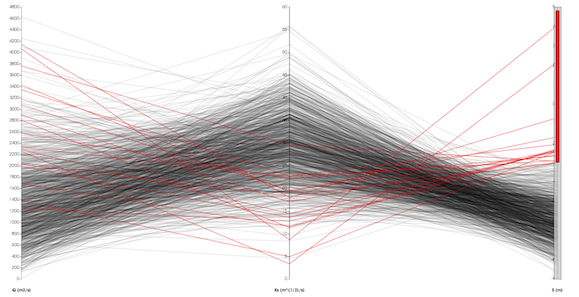
Visualizing the worlds largest simulations introduces a multitude of pragmatic issues. Remotely visualizing Terascale Data on ASC Purple The visualization shown here displays the computed error metrics for a 13 million-cell unstructured data set.ĭistance Visualization for Terascale Data Client side scripting also facilitates automated post-processing verification and ParaView’s multiview capability facilitates comparison of results. The embedded python interpreter allows users to easily build custom parallel computations for analytical solutions, volume integrals, and error metrics over large meshes. Analysts at Sandia National Laboratories are leveraging the python scripting, scalable parallel processing, and multiview technologies of ParaView to simplify the verification of ALEGRA simulations for High Energy Density Physics (HEDP). Verification becomes more difficult with the growth of size and complexity in the simulations. However, simulation results are worthless without verification, which ensures that a simulation can reliably compute the results of a simulation with a known analytical solution. Simulations provide more control over variables and more detail in their outputs than their experimental counterparts while allowing for more runs at a fraction of the price. The rendered field is nut.įor more detailed information refer to the nerdier official ParaView Wiki.Using ParaView to compare simulated and analytical resultsĬomputer simulation is a key component in modern scientific analysis. 
It can be enabled by checking Use FXAA in the Render View section of ParaView’s general settings. Major highlights in this release include: New Rendering Features Anti-aliasing is now available in ParaView. The full list of issues addressed by this release is available here. VolView is another program that provides a simple interface to VTKs 3D scalar field (volume) visualization components. ParaView 5.2.0 is now available for download. Here’s a big big domain rendered on all my cores: Pump case with 6 million cells, rendered with Paraview, on multiple cores. ParaView is an application that allows users to graphically load data sets and visualize them with a commonly used subset of VTK components. You must use CopyStructure before calling this method. Otherwise you will get the error: Structure does not match. Now you can open a case as usual but you must read the decomposed times. Now you open ParaView and instead of opening a file directly, you connect to your server: File > Connect… If you haven’t done it yet, Add Server: Name: your choice C:\Program Files\ParaView\bin>mpiexec -np 4 pvserver In my case, I decompose my cases into 4 subdomains. Also, on Windows Microsoft MPI is run with mpiexec, on Linux mpirun. The number of MPI processes should be equal to number of decomposed subdomains of the OpenFOAM case you will open. So you need to run a server first but with MPI. ParaView is deployed on ASC Purple 'out-of-the-box' despite the complexity of hardware, and ParaViews quality parallel rendering and image delivery mechanism make remotely interacting with the data simple and effective. Rendering results in ParaView on multiple cores on a desktop computer is a little less explained on the light side of the internets. ParaView provides the remote analysis capability our scientists need. Using OpenFOAM in parallel with MPI is well explained in the OpenFOAM manual. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
